Introduction to Data Analysis for Aquatic Sciences
Fundamental aspects of data acquisition, data wrangling, and visualization
In this exercise we will be following a Software Carpentry Lesson. It is suggested that you have this page open to follow along and a bash shell window.
credit:

Copyright (c) Software Carpentry
see also
http://swcarpentry.github.io/shell-novice/
Learning Objectives
- Explain the similarities and differences between a file and a directory.
- Translate an absolute path into a relative path and vice versa.
- Construct absolute and relative paths that identify specific files and directories.
- Identify the actual command, flags, and filenames in a command-line call.
- Demonstrate the use of tab completion, and explain its advantages.
You need to download some files to follow this lesson:
Make a new folder in your Desktop called shell-novice. Download shell-novice-data.zip and move the file to this folder. If it’s not unzipped yet, double-click on it to unzip it. You should end up with a new folder called data.
see also http://gannet.fish.washington.edu/seashell/snaps/pretend-computer.zip
The part of the operating system responsible for managing files and directories is called the file system. It organizes our data into files, which hold information, and directories (also called “folders”), which hold files or other directories.
Several commands are frequently used to create, inspect, rename, and delete files and directories. To start exploring them, let’s open a shell window:
$
The dollar sign is a prompt, which shows us that the shell is waiting for input; your shell may show something more elaborate.
Type the command whoami,
then press the Enter key (sometimes marked Return) to send the command to the shell.
The command’s output is the ID of the current user,
i.e.,
it shows us who the shell thinks we are:
$ whoami
yourname
More specifically, when we type whoami the shell:
whoami,Next,
let’s find out where we are by running a command called pwd
(which stands for “print working directory”).
At any moment,
our current working directory
is our current default directory,
i.e.,
the directory that the computer assumes we want to run commands in
unless we explicitly specify something else.
Here,
the computer’s response is /users/nelle,
which is Nelle’s home directory:
$ pwd
/users/nelle
To understand what a “home directory” is,
let’s have a look at how the file system as a whole is organized.
At the top is the root directory
that holds everything else.
We refer to it using a slash character / on its own;
this is the leading slash in /users/nelle.
Inside that directory are several other directories:
bin (which is where some built-in programs are stored),
data (for miscellaneous data files),
users (where users’ personal directories are located),
tmp (for temporary files that don’t need to be stored long-term),
and so on:
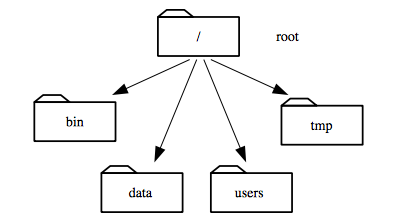
We know that our current working directory /users/nelle is stored inside /users
because /users is the first part of its name.
Similarly,
we know that /users is stored inside the root directory /
because its name begins with /.
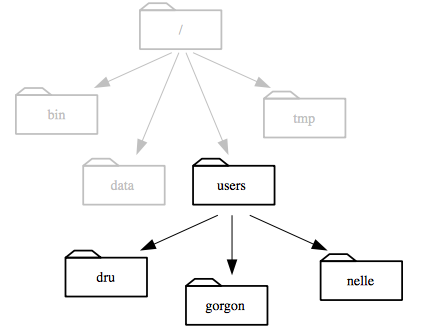
Notice that there are two meanings for the
/character. When it appears at the front of a file or directory name, it refers to the root directory. When it appears inside a name, it’s just a separator.
So that we have the same experience lets navigate to the ‘fake’ directory structure you just downloaded. For example on a Mac you can type cd Desktop/shell-novice/data/users/nelle if you are in your home direcotory.
Let’s see what’s in Nelle’s home directory by running ls,
which stands for “listing”:
$ ls
creatures molecules pizza.cfg
data north-pacific-gyre solar.pdf
Desktop notes.txt writing
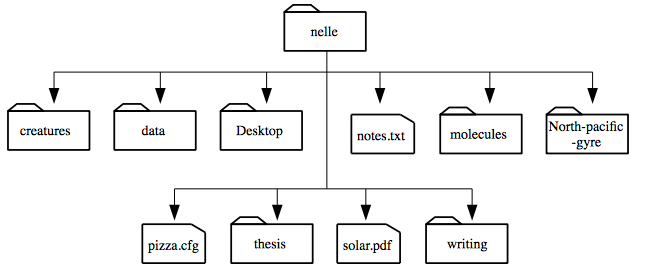
ls prints the names of the files and directories in the current directory in alphabetical order,
arranged neatly into columns.
We can make its output more comprehensible by using the flag -F,
which tells ls to add a trailing / to the names of directories:
$ ls -F
creatures/ molecules/ pizza.cfg
data/ north-pacific-gyre/ solar.pdf
Desktop/ notes.txt writing/
Here,
we can see that /users/nelle contains seven sub-directories.
The names that don’t have trailing slashes,
like notes.txt, pizza.cfg, and solar.pdf,
are plain old files.
And note that there is a space between ls and -F:
without it,
the shell thinks we’re trying to run a command called ls-F,
which doesn’t exist.
Now let’s take a look at what’s in Nelle’s data directory by running ls -F data,
i.e.,
the command ls with the arguments -F and data.
The second argument — the one without a leading dash — tells ls that
we want a listing of something other than our current working directory:
$ ls -F data
amino-acids.txt elements/ morse.txt
pdb/ planets.txt sunspot.txt
The output shows us that there are four text files and two sub-sub-directories. Organizing things hierarchically in this way helps us keep track of our work: it’s possible to put hundreds of files in our home directory, just as it’s possible to pile hundreds of printed papers on our desk, but it’s a self-defeating strategy.
Notice, by the way that we spelled the directory name data.
It doesn’t have a trailing slash:
that’s added to directory names by ls when we use the -F flag to help us tell things apart.
And it doesn’t begin with a slash because it’s a relative path,
i.e., it tells ls how to find something from where we are,
rather than from the root of the file system.
If we run ls -F /data (with a leading slash) we get a different answer,
because /data is an absolute path:
$ ls -F /data
access.log backup/ hardware.cfg
network.cfg
Note you will get an “No file” warning here. This is because we are not really on Nelle’s machine. But this should make sense if you consider the directory organization.
The leading / ( or possibly /c/ if you are using Git Bash on Windows) tells the computer to follow the path from the root of the filesystem,
so it always refers to exactly one directory,
no matter where we are when we run the command.
What if we want to change our current working directory?
Before we do this,
pwd shows us that we’re in /users/nelle,
and ls without any arguments shows us that directory’s contents:
$ pwd
/users/nelle
$ ls
creatures molecules pizza.cfg
data north-pacific-gyre solar.pdf
Desktop notes.txt writing
We can use cd followed by a directory name to change our working directory.
cd stands for “change directory”,
which is a bit misleading:
the command doesn’t change the directory,
it changes the shell’s idea of what directory we are in.
$ cd data
cd doesn’t print anything,
but if we run pwd after it, we can see that we are now in /users/nelle/data.
If we run ls without arguments now,
it lists the contents of /users/nelle/data,
because that’s where we now are:
$ pwd
/users/nelle/data
$ ls -F
amino-acids.txt elements/ morse.txt
pdb/ planets.txt sunspot.txt
We now know how to go down the directory tree: how do we go up? We could use an absolute path:
$ cd /users/nelle
but it’s almost always simpler to use cd .. to go up one level:
$ pwd
/users/nelle/data
$ cd ..
.. is a special directory name meaning
“the directory containing this one”,
or more succinctly,
the parent of the current directory.
Sure enough,
if we run pwd after running cd .., we’re back in /users/nelle:
$ pwd
/users/nelle
The special directory .. doesn’t usually show up when we run ls.
If we want to display it, we can give ls the -a flag:
$ ls -F -a
./ Desktop/ pizza.cfg
../ molecules/ solar.pdf
creatures/ north-pacific-gyre/ writing/
data/ notes.txt
-a stands for “show all”;
it forces ls to show us file and directory names that begin with .,
such as .. (which, if we’re in /users/nelle, refers to the /users directory).
As you can see,
it also displays another special directory that’s just called .,
which means “the current working directory”.
It may seem redundant to have a name for it,
but we’ll see some uses for it soon.
Knowing just this much about files and directories,
Nelle is ready to organize the files that the protein assay machine will create.
First,
she creates a directory called north-pacific-gyre
(to remind herself where the data came from).
Inside that,
she creates a directory called 2012-07-03,
which is the date she started processing the samples.
She used to use names like conference-paper and revised-results,
but she found them hard to understand after a couple of years.
(The final straw was when she found herself creating
a directory called revised-revised-results-3.)
Nelle names her directories “year-month-day”, with leading zeroes for months and days, because the shell displays file and directory names in alphabetical order. If she used month names, December would come before July; if she didn’t use leading zeroes, November (‘11’) would come before July (‘7’).
Each of her physical samples is labelled according to her lab’s convention
with a unique ten-character ID,
such as “NENE01729A”.
This is what she used in her collection log
to record the location, time, depth, and other characteristics of the sample,
so she decides to use it as part of each data file’s name.
Since the assay machine’s output is plain text,
she will call her files NENE01729A.txt, NENE01812A.txt, and so on.
All 1520 files will go into the same directory.
If she is in her home directory, Nelle can see what files she has using the command:
$ ls north-pacific-gyre/2012-07-03/
This is a lot to type, but she can let the shell do most of the work. If she types:
$ ls nor
and then presses tab, the shell automatically completes the directory name for her:
$ ls north-pacific-gyre/
If she presses tab again,
Bash will add 2012-07-03/ to the command,
since it’s the only possible completion.
Pressing tab again does nothing,
since there are 1520 possibilities;
pressing tab twice brings up a list of all the files,
and so on.
This is called tab completion,
and we will see it in many other tools as we go on.
/ on its own is the root directory of the whole filesystem./ on Unix, but \ on Windows... means “the directory above the current one”; . on its own means “the current directory”.something.extension. The extension isn’t required, and doesn’t guarantee anything, but is normally used to indicate the type of data in the file.-.We now know how to explore files and directories, but how do we create them in the first place? Let’s go back to Nelle’s home directory, /users/nelle, and use ls -F to see what it contains:
$ pwd
/users/nelle
Actually will look something more like /Users/sr320/Desktop/shell-novice/data/users/nelle
$ ls -F
creatures/ molecules/ pizza.cfg
data/ north-pacific-gyre/ solar.pdf
Desktop/ notes.txt writing/
Let’s create a new directory called thesis using the command mkdir thesis (which has no output):
$ mkdir thesis
As you might (or might not) guess from its name, mkdir means “make directory”. Since thesis is a relative path (i.e., doesn’t have a leading slash), the new directory is made below the current working directory:
$ ls -F
creatures/ north-pacific-gyre/ thesis/
data/ notes.txt writing/
Desktop/ pizza.cfg
molecules/ solar.pdf
However, there’s nothing in it yet:
$ ls -F thesis
Let’s change our working directory to thesis using cd, then run a text editor called Nano to create a file called draft.txt:
$ cd thesis
$ nano draft.txt
When we say, “nano is a text editor,” we really do mean “text”: it can only work with plain character data, not tables, images, or any other human-friendly media. We use it in examples because almost anyone can drive it anywhere without training, but please use something more powerful for real work. On Unix systems (such as Linux and Mac OS X), many programmers use Emacs or Vim (both of which are completely unintuitive, even by Unix standards), or a graphical editor such as Gedit. On Windows, you may wish to use Notepad++. On a Mac, TextWrangler is an option.
No matter what editor you use, you will need to know where it searches for and saves files. If you start it from the shell, it will (probably) use your current working directory as its default location. If you use your computer’s start menu, it may want to save files in your desktop or documents directory instead. You can change this by navigating to another directory the first time you “Save As…”
Let’s type in a few lines of text, then use Control-O to write our data to disk:
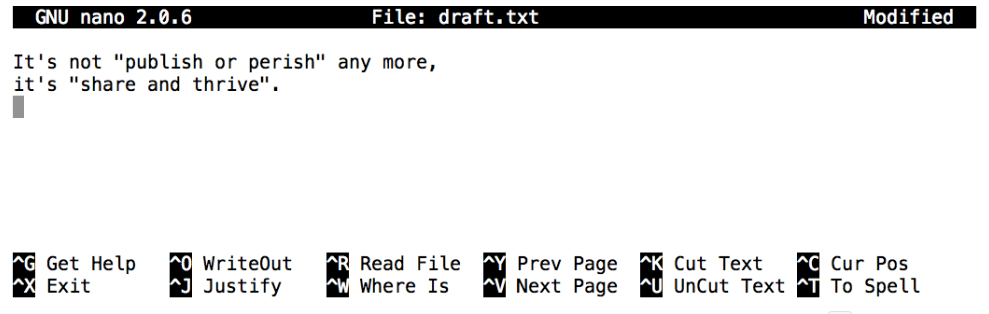
Once our file is saved, we can use Control-X to quit the editor and return to the shell. (Unix documentation often uses the shorthand ^A to mean “control-A”.) nano doesn’t leave any output on the screen after it exits, but ls now shows that we have created a file called draft.txt:
$ ls
draft.txt
Let’s tidy up by running rm draft.txt:
$ rm draft.txt
This command removes files (“rm” is short for “remove”). If we run ls again, its output is empty once more, which tells us that our file is gone:
$ ls
Unix doesn’t have a trash bin: when we delete files, they are unhooked from the file system so that their storage space on disk can be recycled. Tools for finding and recovering deleted files do exist, but there’s no guarantee they’ll work in any particular situation, since the computer may recycle the file’s disk space right away.
Now that we know a few basic commands, we can finally look at the shell’s most powerful feature: the ease with which it lets us combine existing programs in new ways. We’ll start with a directory called molecules that contains six files describing some simple organic molecules. The .pdb extension indicates that these files are in Protein Data Bank format, a simple text format that specifies the type and position of each atom in the molecule.
$ ls molecules
cubane.pdb ethane.pdb methane.pdb
octane.pdb pentane.pdb propane.pdb
Let’s go into that directory with cd and run the command wc *.pdb. wc is the “word count” command: it counts the number of lines, words, and characters in files. The * in *.pdb matches zero or more characters, so the shell turns *.pdb into a complete list of .pdb files:
$ cd molecules
$ wc *.pdb
20 156 1158 cubane.pdb
12 84 622 ethane.pdb
9 57 422 methane.pdb
30 246 1828 octane.pdb
21 165 1226 pentane.pdb
15 111 825 propane.pdb
107 819 6081 total
* is a wildcard. It matches zero or more characters, so *.pdb matches ethane.pdb, propane.pdb, and so on. On the other hand, p*.pdb only matches pentane.pdb and propane.pdb, because the ‘p’ at the front only matches itself.
? is also a wildcard, but it only matches a single character. This means that p?.pdb matches pi.pdb or p5.pdb, but not propane.pdb. We can use any number of wildcards at a time: for example, p*.p?* matches anything that starts with a ‘p’ and ends with ‘.’, ‘p’, and at least one more character (since the ‘?’ has to match one character, and the final * can match any number of characters). Thus, p*.p?* would match preferred.practice, and even p.pi (since the first * can match no characters at all), but not quality.practice (doesn’t start with ‘p’) or preferred.p (there isn’t at least one character after the ‘.p’).
When the shell sees a wildcard, it expands the wildcard to create a list of matching filenames before running the command that was asked for. This means that commands like wc and ls never see the wildcard characters, just what those wildcards matched. This is another example of orthogonal design.
If we run wc -l instead of just wc, the output shows only the number of lines per file:
$ wc -l *.pdb
20 cubane.pdb
12 ethane.pdb
9 methane.pdb
30 octane.pdb
21 pentane.pdb
15 propane.pdb
107 total
We can also use -w to get only the number of words, or -c to get only the number of characters.
Which of these files is shortest? It’s an easy question to answer when there are only six files, but what if there were 6000? Our first step toward a solution is to run the command:
$ wc -l *.pdb > lengths
The > tells the shell to redirect the command’s output to a file instead of printing it to the screen. The shell will create the file if it doesn’t exist, or overwrite the contents of that file if it does. (This is why there is no screen output: everything that wc would have printed has gone into the file lengths instead.) ls lengths confirms that the file exists:
$ ls lengths
lengths
We can now send the content of lengths to the screen using cat lengths. cat stands for “concatenate”: it prints the contents of files one after another. There’s only one file in this case, so cat just shows us what it contains:
$ cat lengths
20 cubane.pdb
12 ethane.pdb
9 methane.pdb
30 octane.pdb
21 pentane.pdb
15 propane.pdb
107 total
Now let’s use the sort command to sort its contents. We will also use the -n flag to specify that the sort is numerical instead of alphabetical. This does not change the file; instead, it sends the sorted result to the screen:
$ sort -n lengths
9 methane.pdb
12 ethane.pdb
15 propane.pdb
20 cubane.pdb
21 pentane.pdb
30 octane.pdb
107 total
We can put the sorted list of lines in another temporary file called sorted-lengths by putting > sorted-lengths after the command, just as we used > lengths to put the output of wc into lengths. Once we’ve done that, we can run another command called head to get the first few lines in sorted-lengths:
$ sort -n lengths > sorted-lengths
$ head -1 sorted-lengths
9 methane.pdb
Using the parameter -1 with head tells it that we only want the first line of the file; -20 would get the first 20, and so on. Since sorted-lengths contains the lengths of our files ordered from least to greatest, the output of head must be the file with the fewest lines.
If you think this is confusing, you’re in good company: even once you understand what wc, sort, and head do, all those intermediate files make it hard to follow what’s going on. We can make it easier to understand by running sort and head together:
$ sort -n lengths | head -1
9 methane.pdb
The vertical bar between the two commands is called a pipe. It tells the shell that we want to use the output of the command on the left as the input to the command on the right. The computer might create a temporary file if it needs to, or copy data from one program to the other in memory, or something else entirely; we don’t have to know or care.
We can use another pipe to send the output of wc directly to sort, which then sends its output to head:
$ wc -l *.pdb | sort -n | head -1
9 methane.pdb
Here’s what actually happens behind the scenes when we create a pipe. When a computer runs a program—any program—it creates a process in memory to hold the program’s software and its current state. Every process has an input channel called standard input. (By this point, you may be surprised that the name is so memorable, but don’t worry: most Unix programmers call it “stdin”. Every process also has a default output channel called standard output (or “stdout”).
The shell is actually just another program. Under normal circumstances, whatever we type on the keyboard is sent to the shell on its standard input, and whatever it produces on standard output is displayed on our screen. When we tell the shell to run a program, it creates a new process and temporarily sends whatever we type on our keyboard to that process’s standard input, and whatever the process sends to standard output to the screen.
Here’s what happens when we run
wc -l *.pdb > lengths. The shell starts by telling the computer to create a new process to run thewcprogram. Since we’ve provided some filenames as parameters,wcreads from them instead of from standard input. And since we’ve used>to redirect output to a file, the shell connects the process’s standard output to that file.If we run
wc -l *.pdb | sort -ninstead, the shell creates two processes (one for each process in the pipe) so thatwcandsortrun simultaneously. The standard output ofwcis fed directly to the standard input ofsort; since there’s no redirection with>,sort’s output goes to the screen. And if we runwc -l *.pdb | sort -n | head -1, we get three processes with data flowing from the files, throughwctosort, and fromsortthroughheadto the screen.This simple idea is why Unix has been so successful. Instead of creating enormous programs that try to do many different things, Unix programmers focus on creating lots of simple tools that each do one job well, and that work well with each other. This programming model is called pipes and filters. We’ve already seen pipes; a filter is a program like
wcorsortthat transforms a stream of input into a stream of output. Almost all of the standard Unix tools can work this way: unless told to do otherwise, they read from standard input, do something with what they’ve read, and write to standard output.The key is that any program that reads lines of text from standard input and writes lines of text to standard output can be combined with every other program that behaves this way as well. You can and should write your programs this way so that you and other people can put those programs into pipes to multiply their power.
As well as using > to redirect a program’s output, we can use < to redirect its input, i.e., to read from a file instead of from standard input. For example, instead of writing wc ammonia.pdb, we could write wc < ammonia.pdb. In the first case, wc gets a command line parameter telling it what file to open. In the second, wc doesn’t have any command line parameters, so it reads from standard input, but we have told the shell to send the contents of ammonia.pdb to wc’s standard input.
Nelle has run her samples through the assay machines and created 1520 files in the north-pacific-gyre/2012-07-03 directory described earlier. As a quick sanity check, she types:
$ cd north-pacific-gyre/2012-07-03
$ wc -l *.txt
The output is 1520 lines that look like this:
300 NENE01729A.txt
300 NENE01729B.txt
300 NENE01736A.txt
300 NENE01751A.txt
300 NENE01751B.txt
300 NENE01812A.txt
... ...
Now she types this:
$ wc -l *.txt | sort -n | head -5
240 NENE02018B.txt
300 NENE01729A.txt
300 NENE01729B.txt
300 NENE01736A.txt
300 NENE01751A.txt
Whoops: one of the files is 60 lines shorter than the others. When she goes back and checks it, she sees that she did that assay at 8:00 on a Monday morning—someone was probably in using the machine on the weekend, and she forgot to reset it. Before re-running that sample, she checks to see if any files have too much data:
$ wc -l *.txt | sort -n | tail -5
300 NENE02040A.txt
300 NENE02040B.txt
300 NENE02040Z.txt
300 NENE02043A.txt
300 NENE02043B.txt
Those numbers look good—but what’s that ‘Z’ doing there in the third-to-last line? All of her samples should be marked ‘A’ or ‘B’; by convention, her lab uses ‘Z’ to indicate samples with missing information. To find others like it, she does this:
$ ls *Z.txt
NENE01971Z.txt NENE02040Z.txt
Sure enough, when she checks the log on her laptop, there’s no depth recorded for either of those samples. Since it’s too late to get the information any other way, she must exclude those two files from her analysis. She could just delete them using rm, but there are actually some analyses she might do later where depth doesn’t matter, so instead, she’ll just be careful later on to select files using the wildcard expression *[AB].txt. As always, the * matches any number of characters; the expression [AB] matches either an ‘A’ or a ‘B’, so this matches all the valid data files she has.
command > file redirects a command’s output to a file.first | second is a pipeline: the output of the first command is used as the input to the second.