Geoduck Gonad Gene Annotations
After kicking around how to make a very big table with all of the annotations… I finally made some progress.
tldr; Annotation table with SwissProt, Gigaton, Ruphibase, Dheilly-comparison.
- also in excel.
todo: need to add sex specific expression.
I started with a list of all 153982 “contigs”.
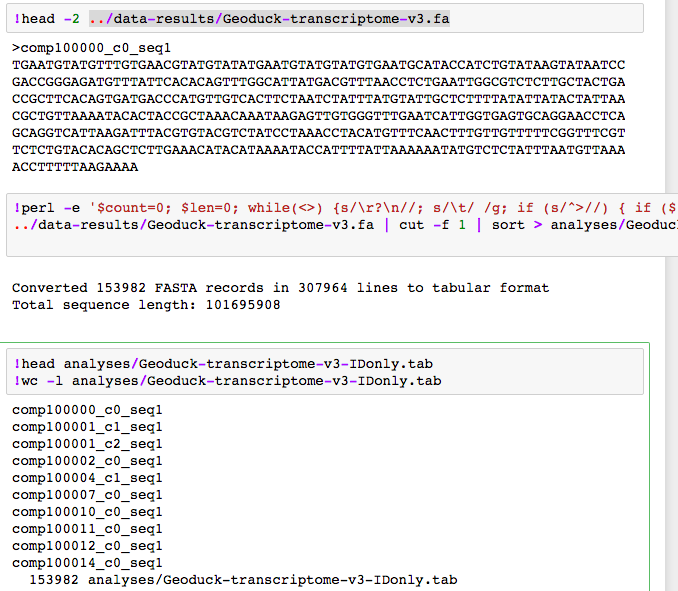
After getting quite fare with join I eventually leveraged SQLShare.
Here are a few bash moments I was fond of…
!join -t $'\t' -a 1 \
analyses/Geoduck-transcriptome-v3-IDonly.tab \
analyses/Geo-v3-join-uniprot-all0916-condensed.sorted \
> analyses/_Master-SP
outputs tab and keeps all of file 1 - important is that -t $'\t' needs to go before arguments.
I cleaned up a blast table
!sort analyses/Geoduck-v3_blastn_GIGAton_e10.out | awk -v OFS='\t' '{ print $1, $11, $2}' \
> analyses/Geoduck-v3_blastn_GIGAton_e10.out.sorted
and with -v OFS='\t' made sure output was tab-delimited.
you can even read in and out on different delimiters…
!awk -F, -v OFS='\t' '{print $2,$11,$13,$14,$20,$21}' analyses/Dheilldy-Gametogenesis-matches.csv
What crushed me was if there was no SP match - the next join went directly adjacent.
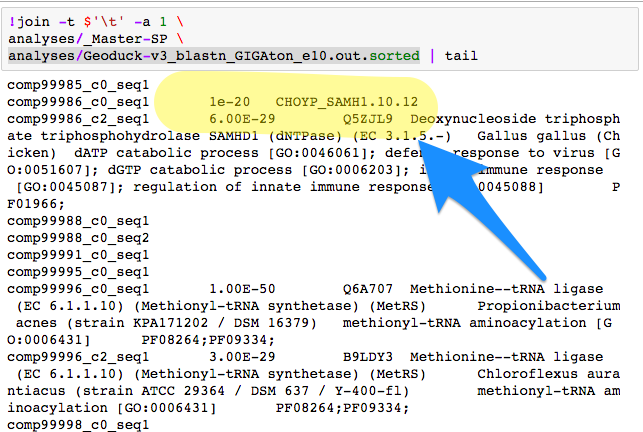
Then I went to SQLShare
The first join was with the SP blast (I could use the bashed up files to upload to SQLShare).
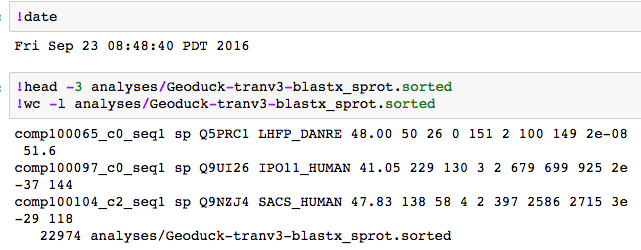
SELECT
*
FROM [roberts.sbr@gmail.com].[Geoduck-transcriptome-v3-IDonly]ID
left join
[roberts.sbr@gmail.com].[Geo-v3-join-uniprot-all0916]sp
on
ID.Column1 = sp.[GeoID]
left join
[roberts.sbr@gmail.com].[Geoduck-v3_blastn_GIGAton_e10]gig
on
ID.Column1 = gig.Column1
left join
[roberts.sbr@gmail.com].[Geoduck-v3_blastn_RuphiBase_e10]ru
on
ID.Column1 = ru.Column1
left join
[roberts.sbr@gmail.com].[Dheilly-comparison-Geo-v3]Dh
on
ID.Column1 = Dh.GEOID
I downloaded this… then !open analyses/_Master-SP-Gig-Ruphi_Dh.csv -a "Microsoft Excel" and succumbed to Excel to clean up a bit and validate.
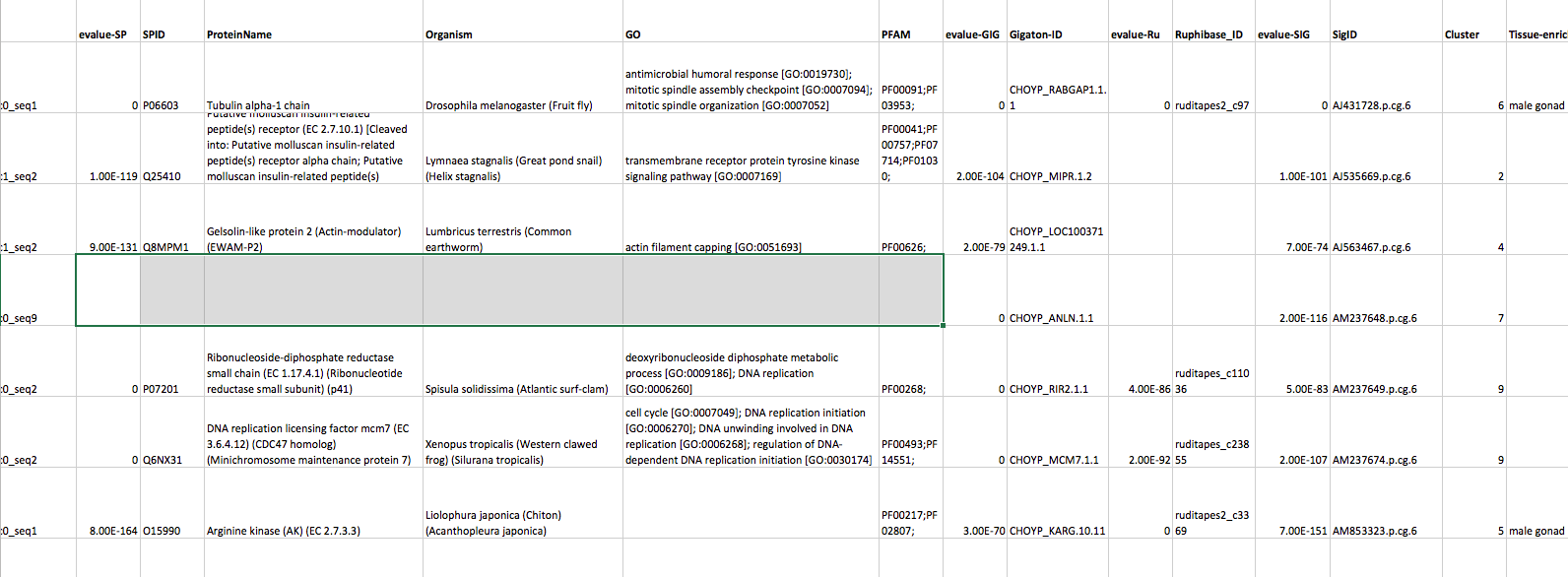
Now each row is a sequence (contig) a 158k row table.
I think all we need now is to add expression data!
